Gene transcriptions/Start sites

The transcription start site is the location where transcription starts at the 5'-end of a gene sequence.[1]
Each human gene is made up of deoxyribonucleic acid (DNA) in a double helix. Along each helix which is composed of a phosphate-deoxyribose polymer are nitrogenous bases. These bases are linked across the helices by hydrogen bonds, one bond per nitrogenous base pair (bp). Each nitrogenous base is part of a nucleotide (nt).
Notations
Notation: let the symbol bp indicate a nitrogenous nucleobase pair as linked by a hydrogen bond between the two antiparallel helices that compose DNA.
Notation: let the symbol nt indicate a nucleotide.
Notation: for the coding of individual nucleobases along a DNA strand, the following letters are standardized by convention:
- adenine - A,
- cytosine - C,
- guanine - G, and
- thymine - T.
Notation: let the subscript (+1) indicate the specific nucleobase along the template strand that is a transcription start site. For example, A+1.
Nitrogenous bases
Nitrogenous bases are typically classified as the derivatives of two parent compounds, pyrimidine and purine.[2]
Three nucleobases found in nucleic acids, cytosine (C), thymine (T), and uracil (U), are pyrimidine derivatives. Of these, cytosine (C) and thymine (T) occur in DNA.
Two of the four bases in nucleic acids, adenine (2) and guanine (3), are purines.
The four nitrogenous bases that occur in DNA linking between the two phosphate-deoxyribose polymer strands are adenine (A), cytosine (C), guanine (G), and thymine (T).
Epigenetics
There are "nearly 50,000 acetylated sites [punctate sites of modified histones] in the human genome that correlate with active transcription start sites and CpG islands and tend to cluster within gene-rich loci."[3]
Gene expressions
Although it is harder to regulate the transcription of genes with multiple transcription start sites, "variations in the expression of a constitutive gene would be minimized by the use of multiple start sites."[4]
Earlier "studies led to the design of a super core promoter (SCP) that contains a TATA, Inr, MTE, and DPE in a single promoter (Juven-Gershon et al., 2006b). The SCP is the strongest core promoter observed in vitro and in cultured cells and yields high levels of transcription in conjunction with transcriptional enhancers. These findings indicate that gene expression levels can be modulated via the core promoter."[4]
Gene transcriptions
"From a teleological standpoint, this arrangement [of focused promoters] is consistent with the notion that it would be easier to regulate the transcription of a gene with a single transcription start site than one with multiple start sites."[4]
Bioinformatics programs usually allow for alternate start codons when searching for protein coding genes.
RNA polymerase II holoenzyme complex may also have to search for one or more transcription start sites.
"RNA polymerase II has been redirected to alternative start sites by reducing ATP concentrations within a nuclear extract, by altering the spacing between the TATA and Inr in a promoter containing both elements, and by dinucleotide initiation strategies".[5]
The core promoter includes the transcription start site(s) (TSS).
RNA polymerase II holoenzyme complexes
RNA polymerase II is recruited to the promoters of protein-coding genes in living cells.[6] Or, transcription factories are present and the euchromatin is brought within the nearest transcription factory and A1BG messenger RNA (mRNA) is transcribed.
For those circumstances in which the holoenzyme is built onto the euchromatin, it is necessary to consider the holoenzyme components and the likely sequence of binding, RNA polymerase II entrance upon the scene and subsequent action.
"RNA polymerase II (also called RNAP II and Pol II) ... catalyzes the transcription of DNA to synthesize precursors of mRNA and most snRNA and microRNA.[7] In humans RNAP II consists of seventeen protein molecules (gene products encoded by POLR2A-L, where the proteins synthesized from 2C-, E-, and F-form homodimers).
Preinitiation complexes

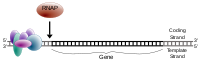
For eukaryotic transcription, the RNA polymerase II holoenzyme de-helicizes the DNA, attaches along the template strand.
Once the preinitiation complex has found its appropriate attachment section along the template strand of DNA, RNA polymerase II is attached and begins transcription.
Start sites
A start site is a biochemically signaled nucleotide or set of nucleotides for attachment either to the Epigenomes or the DNA.
Translation start codon
Specific sequences of DNA act as a template to synthesize mRNA.
The start codon is the first codon of an mRNA transcript translated by a ribosome. The start codon always codes for methionine in eukaryotes
The most common start codon is AUG, ATG in the DNA.
The start codon is almost always preceded by an untranslated region 5' UTR.
For a positive (+) transcription, the start codon on the template strand of DNA is at the end, while a negative (-) transcription has it in the first exon after the 5' UTR.
Alternate start codons (non ATG) are very rare in eukaryote's nuclear genome. Mitochondrial genomes ... use alternate start codons more significantly (mainly GUG and UUG). For example E. coli uses 83% ATG (AUG), 14% GTG (GUG), 3% TTG (UUG) and one or two others (e.g., ATT and CTG).[8]
Note that these alternate start codons are still translated as Met when they are at the start of a protein (even if the codon encodes a different amino acid otherwise). This is because a separate tRNA is used for initiation.
| nonpolar | polar | basic | acidic | (stop codon) |
| 1st base |
2nd base | 3rd base | |||||||
|---|---|---|---|---|---|---|---|---|---|
| T | C | A | G | ||||||
| T | TTT | (Phe/F) Phenylalanine | TCT | (Ser/S) Serine | TAT | (Tyr/Y) Tyrosine | TGT | (Cys/C) Cysteine | T |
| TTC | TCC | TAC | TGC | C | |||||
| TTA | (Leu/L) Leucine | TCA | TAA | Stop (Ochre) | TGA | Stop (Opal) | A | ||
| TTG | TCG | TAG | Stop (Amber) | TGG | (Trp/W) Tryptophan | G | |||
| C | CTT | CCT | (Pro/P) Proline | CAT | (His/H) Histidine | CGT | (Arg/R) Arginine | T | |
| CTC | CCC | CAC | CGC | C | |||||
| CTA | CCA | CAA | (Gln/Q) Glutamine | CGA | A | ||||
| CTG | CCG | CAG | CGG | G | |||||
| A | ATT | (Ile/I) Isoleucine | ACT | (Thr/T) Threonine | AAT | (Asn/N) Asparagine | AGT | (Ser/S) Serine | T |
| ATC | ACC | AAC | AGC | C | |||||
| ATA | ACA | AAA | (Lys/K) Lysine | AGA | (Arg/R) Arginine | A | |||
| ATG[A] | (Met/M) Methionine | ACG | AAG | AGG | G | ||||
| G | GTT | (Val/V) Valine | GCT | (Ala/A) Alanine | GAT | (Asp/D) Aspartic acid | GGT | (Gly/G) Glycine | T |
| GTC | GCC | GAC | GGC | C | |||||
| GTA | GCA | GAA | (Glu/E) Glutamic acid | GGA | A | ||||
| GTG | GCG | GAG | GGG | G | |||||
- The codon ATG both codes for methionine and serves as an initiation site: the first ATG in an mRNA's coding region is where translation into protein begins.[9]
This is the standard or universal genetic code.
This table is found in both DNA Codon Table and Genetic Code. (And probably a few other places.) By default it's the DNA code (using the letter T for Thymine). Use template parameter "T=U" to make it the RNA code (using U for Uracil). See also [Template:Inverse codon table] the inverse codon table.
5'-untranslated region

“The five prime untranslated region (5' UTR), can contain elements for controlling gene expression by way of regulatory elements. It begins at the transcription start site and ends one nucleotide (nt) before the start codon (usually AUG) of the coding region. The 5' UTR has a median length of ~150 nt in eukaryotes, but can be as long as several thousand bases. Several regulatory sequences may be found in the 5' UTR:
- Binding sites for proteins, that may affect the mRNA's stability or translation, for example iron responsive elements, that regulate gene expression in response to iron.
- Riboswitches.
- Sequences that promote or inhibit translation initiation.
- Introns within 5' UTRs have been linked to regulation of gene expression and mRNA export.[10]
Primer extension
Primer extension is a technique whereby the 5' ends of RNA or DNA can be mapped.
"Primer extension can be used to determine the start site of RNA transcription for a known gene."[11]
This technique requires a radiolabelled primer (usually 20 - 50 nucleotides in length) which is complementary to a region near the 3' end of the gene. The primer is allowed to anneal to the RNA and reverse transcriptase is used to synthesize cDNA from the RNA until it reaches the 5' end of the RNA. By running the product on a polyacrylamide gel, it is possible to determine the transcriptional start site, as the length of the sequence on the gel represents the distance from the start site to the radiolabelled primer.
A radioisotope used for primer labeling is 32P.[11]
Focused promoters
"In focused transcription, there is either a single major transcription start site or several start sites within a narrow region of several nucleotides. Focused transcription is the predominant mode of transcription in simpler organisms."[4]
"Focused transcription initiation occurs in all organisms, and appears to be the predominant or exclusive mode of transcription in simpler organisms."[4]
"In vertebrates, focused transcription tends to be associated with regulated promoters".[4]
"The analysis of focused core promoters has led to the discovery of sequence motifs such as the TATA box, BREu (upstream TFIIBrecognition element), Inr (initiator), MTE (motif ten element), DPE (downstream promoter element), DCE (downstream core element), and XCPE1 (Xcore promoter element 1) [...]."[4]
Dispersed promoters
"In dispersed transcription, there are several weak transcription start sites over a broad region of about 50 to 100 nucleotides. Dispersed transcription is the most common mode of transcription in vertebrates. For instance, dispersed transcription is observed in about two-thirds of human genes."[4]
In vertebrates, "dispersed transcription is typically observed in constitutive promoters in CpG islands."[4]
Positions
Positions of nucleotides in the promoter are designated relative to the transcription start site (TSS).
Positions upstream from the TSS are negative numbers counting back from -1, for example, -100 is a position 100 nucleotides upstream from the TSS.
Core promoters

"Focused transcription typically initiates within the Inr, and the A nucleotide in the Inr consensus is usually designed as the “+ 1” position, whether or not transcription actually initiates at that particular nucleotide. This convention is useful because other core promoter motifs, such as the MTE and DPE, function with the Inr in a manner that exhibits a strict spacing dependence with the Inr consensus sequence (and hence, the A + 1 nucleotide) rather than the actual transcription start site (Burke and Kadonaga, 1997, Kutach and Kadonaga, 2000 and Lim et al., 2004)."[4]
"With TATA-driven core promoters, transcription can be achieved in vitro with purified RNA polymerase II, TFIIB, TFIID, TFIIE, TFIIF, and TFIIH."[4]
"NC2 (negative cofactor 2; also known as Dr1-Drap1) [...] was identified as repressor of TATA-dependent transcription [...]."[4]
"TBP (TATA box-binding protein) activates TATA transcription [...] The TBP subunit binds to the TATA box [...] TFIIA appears to promote the binding of TBP to the TATA box."[4]
The core promoter is the minimal portion of the promoter required to properly initiate transcription. The core promoter is approximately -34 nt upstream from the TSS.
The TATA box and the initiator element act synergistically when separated by 25-30 nt but act independently when separated by more than 30 nt.[13] When separated by 15 or 20 bp synergy is retained, and the location of the TSS is dictated by the location of the TATA box rather than the location of the Inr.[13]
GC content
CpG islands typically occur at or near the transcription start site of genes, particularly housekeeping genes, in vertebrates.[14]
GC boxes
"In promoters containing multiple GC boxes but lacking the TATAA box, transcription start sites may be single and specific, as observed in the nerve growth factor receptor gene (42) and the cellular retinol-binding protein gene (37), or there may be multiple heterogeneous start sites, such as those found in the c-myb (4), insulin receptor (45), and Ha-ras (21) genes. ... GC boxes are responsible for directing transcription from the major and the minor start sites. ... All TATAA-less promoters have at least two GC boxes".[15]
"A large subclass of polymerase II promoters lacks both TATAA and CCAAT sequence motifs but contains multiple GC boxes. This promoter class includes several housekeeping genes (e.g., the genes encoding dihydrofolate reductase [DHFR] ..., hydroxymethylglutaryl coenzyme A reductase [39], hypoxanthine guanine phosphoribosyltransferase [33], and adenosine deaminase [46]) [and] nonhousekeeping genes (e.g., the transforming growth factor alpha [9, 23], rat malic enzyme [36], human c-Ha-ras [21], epidermal growth factor receptor [22], and nerve growth factor receptor [42] genes)."[15]
GAAC elements
The GAAC element is usually a core promoter element containing guanine (G), adenine (A), and cytosine (C), "able to direct a new transcription start site 2-7 bases downstream of itself, independent of TATA and Inr regions."[16]
Downstream promoters
"[N]onredundant human promoter sequences 600 bp long (−499 to +100 bp around the TSS) [are available] from [the] Eukaryotic Promoter Database (EPD) release 75 (4, 68) (http://www.epd.isb-sib.ch/), and ... promoters sequences 1,200 bp long (−1,000 to +200 bp) [are available] from the Database of Transcriptional Start Sites (DBTSS) (59, 74, 75) (http://dbtss.hgc.jp/index.html)".[17]
Downstream TFIIB recognition
The downstream TFIIB recognition element (dBRE) has a consensus sequence in the transcription direction on the template strand of 3'-RTDKKKK-5', using degenerate nucleotides, or 3'-A/G-T-A/G/T-G/T-G/T-G/T-G/T-5'.[18]
dBRE is cis-TATA box, between the TATA box and the Inr or transcription start site (TSS) and trans-TSS.[18]
Hypotheses
- RNA polymerase II holoenzyme complex looks for or is keyed by DNA nucleotide sequences to the TSS(s).
- Transcription can be initiated from the template strand, the coding strand, the positive direction and the negative direction.
See also
References
- ↑ Marketa Zvelebil, Jeremy O. Baum (2008). Understanding bioinformatics. Garland Science. ISBN 978-0815340249.
- ↑ Nelson, David L. and Michael M. Cox (2008). Principles of Biochemstry, ed. 5, W.H. Freeman and Company, p. 272.
- ↑ Bradley E. Bernstein, Alexander Meissner, Eric S. Lander (February 23, 2007). "The Mammalian Epigenome". Cell 128 (4): 669–81. doi:10.1016/j.cell.2007.01.033. http://www.sciencedirect.com/science/article/pii/S0092867407001286. Retrieved 19 December 2011.
- ↑ 4.00 4.01 4.02 4.03 4.04 4.05 4.06 4.07 4.08 4.09 4.10 4.11 4.12 Tamar Juven-Gershon and James T. Kadonaga (15 March 2010). "Regulation of gene expression via the core promoter and the basal transcriptional machinery". Developmental Biology 339 (2): 225-9. doi:10.1016/j.ydbio.2009.08.009. http://www.sciencedirect.com/science/article/pii/S0012160609011166. Retrieved 2016-01-16.
- ↑ Stephen T. Smale and James T. Kadonaga (July 2003). "The RNA Polymerase II Core Promoter". Annual Review of Biochemistry 72 (1): 449-79. doi:10.1146/annurev.biochem.72.121801.161520. PMID 12651739. http://www.annualreviews.org/doi/abs/10.1146/annurev.biochem.72.121801.161520. Retrieved 2012-05-07.
- ↑ Myer VE, Young RA (October 1998). "RNA polymerase II holoenzymes and subcomplexes". J. Biol. Chem. 273 (43): 27757–60. doi:10.1074/jbc.273.43.27757. PMID 9774381. http://www.jbc.org/cgi/reprint/273/43/27757.pdf.
- ↑ Kornberg R (1999). "Eukaryotic transcriptional control". Trends in Cell Biology 9 (12): M46. doi:10.1016/S0962-8924(99)01679-7. PMID 10611681.
- ↑ Frederick R. Blattner, Guy Plunkett III, Craig A. Bloch, Nicole T. Perna, Valerie Burland, Monica Riley, Julio Collado-Vides, Jeremy D. Glasner, Christopher K. Rode, George F. Mayhew, Jason Gregor, Nelson Wayne Davis, Heather A. Kirkpatrick, Michael A. Goeden, Debra J. Rose, Bob Mau, Ying Shao (September 1997). "The Complete Genome Sequence of Escherichia coli K-12". Science 277 (5331): 1453-62. doi:10.1126/science.277.5331.1453. http://www.sciencemag.org/content/277/5331/1453.
- ↑ Nakamoto T (March 2009). "Evolution and the universality of the mechanism of initiation of protein synthesis". Gene 432 (1–2): 1–6. doi:10.1016/j.gene.2008.11.001. PMID 19056476.
- ↑ Cenik, C, et al. (2011). "Genome analysis reveals interplay between 5' UTR introns and nuclear mRNA export for secretory and mitochondrial genes". PLoS Genetics 7 (4). doi:10.1371/journal.pgen.1001366. http://www.plosgenetics.org/article/info:doi/10.1371/journal.pgen.1001366.
- ↑ 11.0 11.1 John W. Little (August 18, 2010). Primer extension. Tucson, Arizona USA: University of Arizona. http://www.biochem.arizona.edu/classes/bioc568/primer_extension.htm. Retrieved 2013-02-14.
- ↑ Jennifer E.F. Butler, James T. Kadonaga (October 15, 2002). "The RNA polymerase II core promoter: a key component in the regulation of gene expression". Genes & Development 16 (20): 2583–292. doi:10.1101/gad.1026202. PMID 12381658. http://genesdev.cshlp.org/content/16/20/2583.full.
- ↑ 13.0 13.1 A O'Shea-Greenfield, S T Smale (January 15, 1992). "Roles of TATA and initiator elements in determining the start site location and direction of RNA polymerase II transcription". Journal of Biological Chemistry 267 (2): 1391-402. PMID 1730658. http://www.jbc.org/content/267/2/1391.full.pdf+html. Retrieved 2012-05-16.
- ↑ Saxonov S, Berg P, Brutlag DL (2006). "A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters". Proc Natl Acad Sci USA 103 (5): 1412–1417. doi:10.1073/pnas.0510310103. PMID 16432200. PMC 1345710. //www.ncbi.nlm.nih.gov/pmc/articles/PMC1345710/.
- ↑ 15.0 15.1 Michael C. Blake, Robert C. Jambou, Andrew G. Swick, Jeanne W. Kahn, and Jane Clifford Azizkhan (December 1990). "Transcriptional Initiation Is Controlled by Upstream GC-Box Interactions in a TATAA-Less Promoter". Molecular and Cellular Biology 10 (12): 6632-41. doi:10.1128/MCB.10.12.6632. PMID 2247077. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC362941/pdf/molcellb00048-0544.pdf. Retrieved 2013-01-27.
- ↑ Upinder Singh, Joshua B. Rogers (August 21, 1998). "The Novel Core Promoter Element GAAC in the hgl5 Gene of Entamoeba histolytica Is Able to Direct a Transcription Start Site Independent of TATA or Initiator Regions". The Journal of Biological Chemistry 273 (34): 21663-8. doi:10.1074/jbc.273.34.21663. http://www.jbc.org/content/273/34/21663.full. Retrieved 2013-02-13.
- ↑ Dong-Hoon Lee, Naum Gershenzon, Malavika Gupta, Ilya P. Ioshikhes, Danny Reinberg and Brian A. Lewis (November 2005). "Functional Characterization of Core Promoter Elements: the Downstream Core Element Is Recognized by TAF1". Molecular and Cellular Biology 25 (21): 9674-86. doi:10.1128/MCB.25.21.9674-9686.2005. PMID 16227614. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1265815/. Retrieved 2010-10-23.
- ↑ 18.0 18.1 Wensheng Deng, Stefan G.E. Roberts (October 15, 2005). "A core promoter element downstream of the TATA box that is recognized by TFIIB". Genes & Development 19 (20): 2418–23. doi:10.1101/gad.342405. PMID 16230532. http://genesdev.cshlp.org/content/19/20/2418.full.
Further reading
- Javahery R, Khachi A, Lo K, Zenzie-Gregory B, Smale ST (January 1994). "DNA Sequence Requirements for Transcriptional Initiator Activity in Mammalian Cells". Mol Cell Biol. 14 (1): 116-27. PMID 8264580. http://mcb.asm.org/cgi/reprint/14/1/116?view=long&pmid=8264580.
- Liston DR, Johnson PJ (March 1999). "Analysis of a Ubiquitous Promoter Element in a Primitive Eukaryote: Early Evolution of the Initiator Element". Mol Cell Biol. 19 (3): 2380-8. http://mcb.asm.org/cgi/content/full/19/3/2380.
- Myer VE, Young RA (October 1998). "RNA polymerase II holoenzymes and subcomplexes". J. Biol. Chem. 273 (43): 27757–60. doi:10.1074/jbc.273.43.27757. PMID 9774381. http://www.jbc.org/cgi/reprint/273/43/27757.pdf.
- Tsvetkov S, Ivanova E, Djondjurov L (December 1989). "The pool of histones in the nucleosol and cytosol of proliferating Friend cells is small, uneven and chasable". Biochem J. 264 (3): 785–91. PMID 2619716. PMC 1133654. //www.ncbi.nlm.nih.gov/pmc/articles/PMC1133654/.
External links
- African Journals Online
- Bing Advanced search
- GenomeNet KEGG database
- Google Books
- Google scholar Advanced Scholar Search
- Home - Gene - NCBI
- JSTOR
- Lycos search
- NCBI All Databases Search
- NCBI Site Search
- PubChem Public Chemical Database
- Questia - The Online Library of Books and Journals
- SAGE journals online
- Scirus for scientific information only advanced search
- SpringerLink
- Taylor & Francis Online
- WikiDoc The Living Textbook of Medicine
- Wiley Online Library Advanced Search
- Yahoo Advanced Web Search
