Gene transcriptions/Elements/Factor II B recognitions
The B recognition element (BRE) is a DNA sequence found in the promoter region of most genes in eukaryotes and Archaea.[1][2]
The BRE is a cis-regulatory element that is found immediately upstream of the TATA box, and consists of 7 nucleotides.
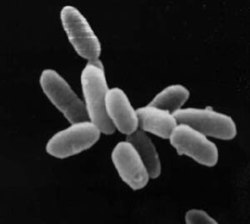
In the archaean from the Dead Sea imaged at the right, "We have completely fragmented their DNA. I mean we have completely destroyed it by bombarding it with [radiation]. And they can reassemble their entire chromosome and put it back into working order within several hours."[3]
Consensus sequences
[edit | edit source]
The consensus sequence is 5’-G/C G/C G/A C G C C-3’.[4]
The general consensus sequence using degenerate nucleotides is 5’-SSRCGCC-3’, where S = G or C and R = A or G.[5]
Transcription start sites
[edit | edit source]
"The position in nucleotides (nt) relative to the transcription start site (TSS, +1)" is -35 for the BRE. Of human promoters, some "22-25% [are] BRE containing promoters ... the functional consensus sequences for BRE ... motif [is] still poorly defined."[5]
General transcription factor II Bs
[edit | edit source]The Transcription Factor IIB (TFIIB) recognizes this sequence in the DNA, and binds to it. The fourth and fifth alpha helices of TFIIB intercalate with the major groove of the DNA at the BRE. TFIIB is one part of the preinitiation complex that helps RNA Polymerase II bind to the DNA.
Core promoters
[edit | edit source]
The core promoter is approximately -34 nts upstream from the TSS.
From the first nucleotide just after ZSCAN22 to the first nucleotide just before A1BG are 4460 nucleotides. The core promoter on this side of A1BG extends from approximately 4425 to the possible transcription start site at nucleotide number 4460.
To extend the analysis from inside and just on the other side of ZNF497 some 3340 nts have been added to the data. This would place the core promoter some 3340 nts further away from the other side of ZNF497. The TSS would be at about 4300 nts with the core promoter starting at 4266.
Def. "the factors, including RNA polymerase II itself, that are minimally essential for transcription in vitro from an isolated core promoter" is called the basal machinery, or basal transcription machinery.[7]
Hypotheses
[edit | edit source]- B recognition factor is not involved in the transcription of A1BG.
- If involved it assists transcription by other TFs.
Samplings
[edit | edit source]Regarding hypothesis 1
[edit | edit source]The B recognition element (BRE) is not involved in the transcription of A1BG.
For the Basic programs (starting with SuccessablesBRE.bas) written to compare nucleotide sequences with the sequences on either the template strand (-), or coding strand (+), of the DNA, in the negative direction (-), or the positive direction (+), the programs are, are looking for, and found:
- negative strand in the negative direction (from ZSCAN22 to A1BG) is SuccessablesBRE--.bas, looking for 3'-(G/C)(G/C)(G/A)CGCC-5', 3, 3'-CCACGCC-5' at 380, 3'-CCGCGCC-5' at 1762, and 3'-CCACGCC-5' at 2197,
- negative strand in the positive direction (from ZNF497 to A1BG) is SuccessablesBRE-+.bas, looking for 3'-(G/C)(G/C)(G/A)CGCC-5', 3, 3'-GCACGCC-5', 1302, 3'-GGACGCC-5', 1672, 3'-GGGCGCC-5', 1769,
- positive strand in the negative direction is SuccessablesBRE+-.bas, looking for 3'-(G/C)(G/C)(G/A)CGCC-5', 0,
- positive strand in the positive direction is SuccessablesBRE++.bas, looking for 3'-(G/C)(G/C)(G/A)CGCC-5', 3, 3'-CCACGCC-5', 489, 3'-CGACGCC-5', 1033, 3'-CCACGCC-5', 1764,
- complement, negative strand, negative direction is SuccessablesBREc--.bas, looking for 3'-G/C-G/C-C/T-G-C-G-G-5', 1, 3'-CCTGCGG-5' at 1153,
- complement, negative strand, positive direction is SuccessablesBREc-+.bas, looking for 3'-G/C-G/C-C/T-G-C-G-G-5', 3, 3'-GGTGCGG-5', 489, 3'-GCTGCGG-5', 1033, 3'-GGTGCGG-5', 1764,
- complement, positive strand, negative direction is SuccessablesBREc+-.bas, looking for 3'-G/C-G/C-C/T-G-C-G-G-5', 3, 3'-GGTGCGG-5' at 380, 3'-GGCGCGG-5' at 1762, and 3'-GGTGCGG-5' at 2197,
- complement, positive strand, negative direction is SuccessablesBREc++.bas, looking for 3'-G/C-G/C-C/T-G-C-G-G-5', 3, 3'-CGTGCGG-5', 1302, 3'-CCTGCGG-5', 1672, 3'-CCCGCGG-5', 1769,
- inverse complement, negative strand, negative direction is SuccessablesBREci--.bas, looking for 3'-G-G-C-G-C/T-G/C-G/C-5', 0,
- inverse complement, negative strand, positive direction is SuccessablesBREci-+.bas, looking for 3'-G-G-C-G-C/T-G/C-G/C-5', 1, 3'-GGCGCCC-5', 1770,
- inverse complement, positive strand, negative direction is SuccessablesBREci+-.bas, looking for 3'-G-G-C-G-C/T-G/C-G/C-5', 4, 3'-GGCGTGG-5' at 1244, 3'-GGCGCGG-5' at 1762, 3'-GGCGTGG-5' at 1897, and 3'-GGCGTGG-5' at 3047,
- inverse complement, positive strand, positive direction is SuccessablesBREci++.bas, looking for 3'-G-G-C-G-C/T-G/C-G/C-5', 4, 3'-GGCGCGC-5', 682, 3'-GGCGCCG-5', 1338, 3'-GGCGCCG-5', 1438, 3'-GGCGTGG-5', 2566,
- inverse, negative strand, negative direction, is SuccessablesBREi--.bas, looking for 3'-C-C-G-C-G/A-G/C-G/C-5', 4, 3'-CCGCACC-5' at 1244, 3'-CCGCGCC-5' at 1762, 3'-CCGCACC-5' at 1897, and 3'-CCGCACC-5' at 3047,
- inverse, negative strand, positive direction, is SuccessablesBREi-+.bas, looking for 3'-C-C-G-C-G/A-G/C-G/C-5', 4, 3'-CCGCGCG-5', 682, 3'-CCGCGGC-5', 1338, 3'-CCGCGGC-5', 1438, 3'-CCGCACC-5', 2566,
- inverse, positive strand, negative direction, is SuccessablesBREi+-.bas, looking for 3'-C-C-G-C-G/A-G/C-G/C-5', 0,
- inverse, positive strand, positive direction, is SuccessablesBREi++.bas, looking for 3'-C-C-G-C-G/A-G/C-G/C-5', 1, 3'-CCGCGGG-5', 1770.
Regarding hypothesis 2
[edit | edit source]
The BRE is likely involved in and assists transcription by these other TFs when they are present.
The diagram on the right shows an overview of the four core promoter elements: B recognition element (BRE), TATA box, initiator element (Inr), and downstream promoter element (DPE), with their respective consensus sequences and their distance from the transcription start site.[6]
On the left is a more comprehensive diagram of a promoter. "The best known core promoter element is the TATA-box, consisting of an AT-rich sequence located ~27 bp upstream of the TSS, but several other core promoter elements exist, including initiator element (Inr) and X core promoter element 1 (XCPE1) localized around the TSS, the TFIIB recognition elements (BRE) that are positioned upstream of the TSS, and downstream promoter element (DPE), motif ten element (MTE) and downstream core element (DCE) that are situated downstream of TSS. The distal regulatory elements include locus control regions (LCR), enhancers, silencers and insulators. The enhancers and silencers have sites for binding multiple transcription factors and they function in activating and repressing transcription, respectively. Insulators operate by blocking genes from being affected by the regulatory elements of neighbouring genes. The LCR consists of multiple transcription regulatory elements that function together to provide proper expression regulation to a cluster of genes."[8]
Transcribed BREs
[edit | edit source]"One of the major discoveries in large-scale detection of promoters was the existence of different classes of core promoters, for which there are common features across the metazoan lineage. The number of main classes has not been settled, but the current evidence points towards three main functional classes [...]."[9]
"In D. melanogaster, a number of different promoter types have been suggested based on motif content. An exhaustive analysis of motif composition in D. melanogaster and human promoters14 revealed extensive differences in the type and directionality of motifs found in different promoters and their association with gene function. In parallel, five principal motif-based classes of D. melanogaster promoters were proposed15, which could be further grouped into three general functional classes16."[9]
"Type I consists of the tissue-specific promoters, which are similar to the low-CpG class in mammals with respect to motif composition, stage of development at which they are expressed and tissue specificity, and they are characterized by a high enrichment for a TATA box at an appropriate distance from an initiator element (Inr element). Type II promoters are associated with ‘housekeeping’ genes and genes that are regulated at the level of individual cells; they have either a DNA recognition element (DRE) or a combination of novel motifs15. Finally, type III promoters have an Inr element only or an Inr element plus a downstream promoter element (DPE). These promoters are preferentially associated with developmentally regulated genes, the expression of which is precisely coordinated across different cells in a tissue or anatomical structure16."[9]
See also
[edit | edit source]References
[edit | edit source]- ↑ Lagrange T, Kapanidis AN, Tang H, Reinberg D, Ebright RH (1998). "New core promoter element in RNA polymerase II-dependent transcription: sequence-specific DNA binding by transcription factor IIB". Genes & Development 12 (1): 34–44. doi:10.1101/gad.12.1.34. PMID 9420329. PMC 316406. //www.ncbi.nlm.nih.gov/pmc/articles/PMC316406/.
- ↑ Littlefield O, Korkhin Y, Sigler PB (1999). "The structural basis for the oriented assembly of a TBP/TFB/promoter complex". Proceedings of the National Academy of Sciences of the USA 96 (24): 13668–73. doi:10.1073/pnas.96.24.13668. PMID 10570130. PMC 24122. //www.ncbi.nlm.nih.gov/pmc/articles/PMC24122/.
- ↑ Adrienne Kish (September 10, 2004). Secrets of a Salty Survivor A microbe that grows in the Dead Sea is teaching scientists about the art of DNA repair. Washington, DC USA: NASA. http://science1.nasa.gov/science-news/science-at-nasa/2004/10sep_radmicrobe/. Retrieved 2014-05-15.
- ↑ Alan K. Kutach, James T. Kadonaga (July 2000). "The Downstream Promoter Element DPE Appears To Be as Widely Used as the TATA Box in Drosophila Core Promoters". Molecular and Cellular Biology 20 (13): 4754-64. PMID 10848601. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC85905/pdf/mb004754.pdf. Retrieved 2012-07-15.
- ↑ 5.0 5.1 Chuhu Yang, Eugene Bolotin, Tao Jiang, Frances M. Sladek, Ernest Martinez. (March 7, 2007). "Prevalence of the initiator over the TATA box in human and yeast genes and identification of DNA motifs enriched in human TATA-less core promoters". Gene 389 (1): 52-65. doi:10.1016/j.gene.2006.09.029. PMID 17123746. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1955227/?tool=pubmed.
- ↑ 6.0 6.1 Jennifer E.F. Butler, James T. Kadonaga (October 15, 2002). "The RNA polymerase II core promoter: a key component in the regulation of gene expression". Genes & Development 16 (20): 2583–292. doi:10.1101/gad.1026202. PMID 12381658. http://genesdev.cshlp.org/content/16/20/2583.full.
- ↑ Stephen T. Smale and James T. Kadonaga (July 2003). "The RNA Polymerase II Core Promoter". Annual Review of Biochemistry 72 (1): 449-79. doi:10.1146/annurev.biochem.72.121801.161520. PMID 12651739. http://www.lps.ens.fr/~monasson/Houches/Kadonaga/CorePromoterAnnuRev2003.pdf. Retrieved 2012-05-07.
- ↑ Alexandra Elsing (2014). Regulation of HSF2 and its function in mitosis. Turku, Finland: Department of Biosciences, Åbo Akademi University. pp. 123. ISBN 978-952-12-3105-6. http://www.doria.fi/bitstream/handle/10024/98908/elsing_alexandra.pdf?sequence=2&isAllowed=y. Retrieved 2018-04-22.
- ↑ 9.0 9.1 9.2 Boris Lenhard, Albin Sandelin and Piero Carninci (April 2012). "Metazoan promoters: emerging characteristics and insights into transcriptional regulation". Nature Reviews Genetics 13: 233-245. http://www.academia.edu/download/41359722/REGULATORY_ELEMENTS_Metazoan_promoters_e20160120-1921-11sua5q.pdf. Retrieved 2018-4-30.
Further reading
[edit | edit source]- Chuhu Yang, Eugene Bolotin, Tao Jiang, Frances M. Sladek, Ernest Martinez. (March 7, 2007). "Prevalence of the initiator over the TATA box in human and yeast genes and identification of DNA motifs enriched in human TATA-less core promoters". Gene 389 (1): 52-65. doi:10.1016/j.gene.2006.09.029. PMID 17123746. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC1955227/?tool=pubmed.
External links
[edit | edit source]- African Journals Online
- Bing Advanced search
- GenomeNet KEGG database
- Google Books
- Google scholar Advanced Scholar Search
- Home - Gene - NCBI
- JSTOR
- Lycos search
- NCBI All Databases Search
- Office of Scientific & Technical Information
- PubChem Public Chemical Database
- Questia - The Online Library of Books and Journals
- SAGE journals online
- Scirus for scientific information only advanced search
- SpringerLink
- Taylor & Francis Online
- WikiDoc The Living Textbook of Medicine
- Wiley Online Library Advanced Search
- Yahoo Advanced Web Search
