WikiJournal Preprints/Genome-Wide Association Studies
This article is an unpublished pre-print not yet undergoing peer review.
To submit this article for peer review, please:
Article information
Abstract
In genetics, a genome-wide association study (GWA study, or GWAS), also known as whole genome association study (WGA study, or WGAS), is an observational study of a genome-wide set of genetic variants| in different individuals to see if any variant is associated with a trait. GWASs typically focus on associations between single-nucleotide polymorphisms (SNPs) and traits like major human diseases, but can equally be applied to any other organism.
When applied to human data, GWA studies compare the DNA of participants having varying phenotypes for a particular trait or disease. These participants may be people with a disease (cases) and similar people without the disease (controls), or they may be people with different phenotypes for a particular trait, for example blood pressure. This approach is known as phenotype-first, in which the participants are classified first by their clinical manifestation(s), as opposed to genotype-first|. Each person gives a sample of DNA, from which millions of genetic variants| are read using SNP arrays. If one type of the variant (one allele) is more frequent in people with the disease, the variant is said to be associated with the disease. The associated SNPs are then considered to mark a region of the human genome that may influence the risk of disease.
GWA studies investigate the entire genome, in contrast to methods that specifically test a small number of pre-specified genetic regions. Hence, GWAS is a non-candidate-driven approach, in contrast to gene-specific candidate-driven studies|. GWA studies identify SNPs and other variants in DNA associated with a disease, but they cannot on their own specify which genes are causal.[2][3][4]
The first successful GWAS was published in 2002 studying myocardial infarction.[5] This design study was then implemented in the landmark GWA 2005 study investigating patients with age-related macular degeneration and found two SNPs with significantly altered allele frequency compared to healthy controls.[6] As of 2017[update] , hundreds or thousands of individuals are tested in a typical GWA study. Over 3,000 human GWA studies have examined over 1,800 diseases and traits, and thousands of SNP associations have been found.[7]
Background
[edit | edit source]
Any two human genomes differ in millions of different ways. There are small variations in the individual nucleotides of the genomes (SNPs|) as well as many larger variations, such as deletions|, insertions| and copy number variations. Any of these may cause alterations in an individual's traits, or phenotype, which can be anything from disease risk to physical properties such as height.[9] Around the year 2000, prior to the introduction of GWA studies, the primary method of investigation was through inheritance studies of genetic linkage in families. This approach had proven highly useful towards single gene disorders|.[10]Template:Failed verification However, for common and complex diseases the results of genetic linkage studies proved hard to reproduce.[9][11] A suggested alternative to linkage studies was the genetic association study. This study type asks if the allele of a genetic variant| is found more often than expected in individuals with the phenotype of interest (e.g. with the disease being studied). Early calculations on statistical power indicated that this approach could be better than linkage studies at detecting weak genetic effects.[12]
In addition to the conceptual framework several additional factors enabled the GWA studies. One was the advent of biobanks, which are repositories of human genetic material that greatly reduced the cost and difficulty of collecting sufficient numbers of biological specimens for study.[13] Another was the International HapMap Project, which, from 2003 identified a majority of the common SNPs interrogated in a GWA study.[14] The haploblock structure| identified by HapMap project also allowed the focus on the subset of SNPs that would describe most of the variation. Also the development of the methods to genotype all these SNPs using genotyping arrays| was an important prerequisite.[15]
Methods
[edit | edit source]
The most common approach of GWA studies is the case-control| setup, which compares two large groups of individuals, one healthy control group and one case group affected by a disease. All individuals in each group are genotyped for the majority of common known SNPs. The exact number of SNPs depends on the genotyping technology, but are typically one million or more.[8] For each of these SNPs it is then investigated if the allele frequency is significantly altered between the case and the control group.[17] In such setups, the fundamental unit for reporting effect sizes is the odds ratio. The odds ratio is the ratio of two odds, which in the context of GWA studies are the odds of disease for individuals having a specific allele and the odds of disease for individuals who do not have that same allele. When the allele frequency in the case group is much higher than in the control group, the odds ratio is higher than 1, and vice versa for lower allele frequency. Additionally, a P-value for the significance of the odds ratio is typically calculated using a simple chi-squared test. Finding odds ratios that are significantly different from 1 is the objective of the GWA study because this shows that a SNP is associated with disease.[17]
There are several variations to this case-control approach. A common alternative to case-control GWA studies is the analysis of quantitative phenotypic data, e.g. height or biomarker| concentrations or even gene expression|. Likewise, alternative statistics designed for dominance| or recessive penetrance patterns can be used.[17] Calculations are typically done using bioinformatics software such as SNPTEST and PLINK, which also include support for many of these alternative statistics.[16][18] Earlier GWAS focused on the effect of individual SNPs. However, the empirical evidence shows that complex interactions among two or more SNPs, epistasis, might contribute to complex diseases. Moreover, the researchers try to integrate GWA data with other biological data such as protein protein interaction network| to extract more informative results.[19][20]
A key step in the majority of GWA studies is the imputation| of genotypes at SNPs not on the genotype chip used in the study.[21] This process greatly increases the number of SNPs that can be tested for association, increases the power of the study, and facilitates meta-analysis of GWAS across distinct cohorts. Genotype imputation is carried out by statistical methods that combine the GWAS data together with a reference panel of haplotypes. These methods take advantage of sharing of haplotypes between individuals over short stretches of sequence to impute alleles. Existing software packages for genotype imputation include IMPUTE2[22] and MaCH.[23]
In addition to the calculation of association, it is common to take into account any variables that could potentially confound| the results. Sex and age are common examples of confounding variables. Moreover, it is also known that many genetic variations are associated with the geographical and historical populations in which the mutations first arose.[24] Because of this association, studies must take account of the geographic and ethnic background of participants by controlling for what is called population stratification. If they fail to do so, these studies can produce false positive results.[25]
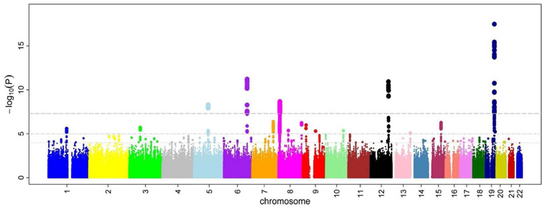
After odds ratios and P-values have been calculated for all SNPs, a common approach is to create a Manhattan plot. In the context of GWA studies, this plot shows the negative logarithm of the P-value as a function of genomic location. Thus the SNPs with the most significant association stand out on the plot, usually as stacks of points because of haploblock structure. Importantly, the P-value threshold for significance is corrected for multiple testing| issues. The exact threshold varies by study,[26] but the conventional threshold is 5×10−8
to be significant in the face of hundreds of thousands to millions of tested SNPs.[8][17][27] GWA studies typically perform the first analysis in a discovery cohort, followed by validation of the most significant SNPs in an independent validation cohort.
Results
[edit | edit source]
Attempts have been made at creating comprehensive catalogues of SNPs that have been identified from GWA studies.[29] As of 2009, SNPs associated with diseases are numbered in the thousands.[30]
The first GWA study, conducted in 2005, compared 96 patients with age-related macular degeneration (ARMD) with 50 healthy controls.[31] It identified two SNPs with significantly altered allele frequency between the two groups. These SNPs were located in the gene encoding complement factor H, which was an unexpected finding in the research of ARMD. The findings from these first GWA studies have subsequently prompted further functional research towards therapeutical manipulation of the complement system in ARMD.[32] Another landmark publication in the history of GWA studies was the Wellcome Trust Case Control Consortium (WTCCC) study, the largest GWA study ever conducted at the time of its publication in 2007. The WTCCC included 14,000 cases of seven common diseases (~2,000 individuals for each of coronary heart disease, type 1 diabetes|, type 2 diabetes|, rheumatoid arthritis, Crohn's disease, bipolar disorder, and hypertension) and 3,000 shared controls.[16] This study was successful in uncovering many new disease genes underlying these diseases.[16][33]
Since these first landmark GWA studies, there have been two general trends.[34] One has been towards larger and larger sample sizes. In 2018, several genome-wide association studies are reaching a total sample size of over 1 million participants, including 1.1 million in a genome-wide study of educational attainment[35] and a study of insomnia containing 1.3 million individuals.[36] The reason is the drive towards reliably detecting risk-SNPs that have smaller odds ratios and lower allele frequency. Another trend has been towards the use of more narrowly defined phenotypes, such as blood lipids, proinsulin or similar biomarkers.[37][38] These are called intermediate phenotypes, and their analyses may be of value to functional research into biomarkers.[39] A variation of GWAS uses participants that are first-degree relatives of people with a disease. This type of study has been named genome-wide association study by proxy (GWAX).[40]
A central point of debate on GWA studies has been that most of the SNP variations found by GWA studies are associated with only a small increased risk of the disease, and have only a small predictive value. The median odds ratio is 1.33 per risk-SNP, with only a few showing odds ratios above 3.0.[2][41] These magnitudes are considered small because they do not explain much of the heritable variation. This heritable| variation is known from heritability studies based on monozygotic twins.[42] For example, it is known that 80-90% of variance in height can be explained by hereditary differences, but GWA studies only account for a minority of this variance.[42]
Clinical applications
[edit | edit source]A challenge for future successful GWA study is to apply the findings in a way that accelerates drug| and diagnostics development, including better integration of genetic studies into the drug-development process and a focus on the role of genetic variation in maintaining health as a blueprint for designing new drugs| and diagnostics|.[43] Several studies have looked into the use of risk-SNP markers as a means of directly improving the accuracy of prognosis. Some have found that the accuracy of prognosis improves,[44] while others report only minor benefits from this use.[45] Generally, a problem with this direct approach is the small magnitudes of the effects observed. A small effect ultimately translates into a poor separation of cases and controls and thus only a small improvement of prognosis accuracy. An alternative application is therefore the potential for GWA studies to elucidate pathophysiology.[46]
One such success is related to identifying the genetic variant associated with response to anti-hepatitis C virus treatment. For genotype 1 hepatitis C treated with Pegylated interferon-alpha-2a or Pegylated interferon-alpha-2b combined with ribavirin, a GWA study[47] has shown that SNPs near the human IL28B| gene, encoding interferon lambda 3, are associated with significant differences in response to the treatment. A later report demonstrated that the same genetic variants are also associated with the natural clearance of the genotype 1 hepatitis C virus.[48] These major findings facilitated the development of personalized medicine and allowed physicians to customize medical decisions based on the patient's genotype.[49]
The goal of elucidating pathophysiology has also led to increased interest in the association between risk-SNPs and the gene expression of nearby genes, the so-called expression quantitative trait loci (eQTL) studies.[50] The reason is that GWAS studies identify risk-SNPs, but not risk-genes, and specification of genes is one step closer towards actionable drug targets|. As a result, major GWA studies by 2011 typically included extensive eQTL analysis.[51][52][53] One of the strongest eQTL effects observed for a GWA-identified risk SNP is the SORT1 locus.[37] Functional follow up studies of this locus using small interfering RNA and gene knock-out mice| have shed light on the metabolism of low-density lipoproteins, which have important clinical implications for cardiovascular disease.[37][54][55]
Limitations
[edit | edit source]GWA studies have several issues and limitations that can be taken care of through proper quality control and study setup. Lack of well defined case and control groups, insufficient sample size, control for multiple testing| and control for population stratification are common problems.[3] Particularly the statistical issue of multiple testing wherein it has been noted that "the GWA approach can be problematic because the massive number of statistical tests performed presents an unprecedented potential for false-positive results".[3] Ignoring these correctible issues has been cited as contributing to a general sense of problems with the GWA methodology.[56] In addition to easily correctible problems such as these, some more subtle but important issues have surfaced. A high-profile GWA study that investigated individuals with very long life spans to identify SNPs associated with longevity is an example of this.[57] The publication came under scrutiny because of a discrepancy between the type of genotyping array| in the case and control group, which caused several SNPs to be falsely highlighted as associated with longevity.[58] The study was subsequently retracted|,[59] but a modified manuscript was later published.[60]
In addition to these preventable issues, GWA studies have attracted more fundamental criticism, mainly because of their assumption that common genetic variation plays a large role in explaining the heritable variation of common disease.[61] This aspect of GWA studies has attracted the criticism that, although it could not have been known prospectively, GWA studies were ultimately not worth the expenditure.[46] Alternative strategies suggested involve linkage analysis. More recently, the rapidly decreasing price of complete genome sequencing have also provided a realistic alternative to genotyping array|-based GWA studies. It can be discussed if the use of this new technique is still referred to as a GWA study, but high-throughput sequencing does have potential to side-step some of the shortcomings of non-sequencing GWA.[62]
Fine-mapping
[edit | edit source]Genotyping arrays designed for GWAS rely on linkage disequilibrium to provide coverage of the entire genome by genotyping a subset of variants. Because of this, the reported associated variants are unlikely to be the actual causal variants. Associated regions can contain hundreds of variants spanning large regions and encompassing many different genes, making the biological interpretation of GWAS loci more difficult. Fine-mapping is a process to refine these lists of associated variants to a credible set most likely to include the causal variant.
Fine-mapping requires all variants in the associated region to have been genotyped or imputed (dense coverage), very stringent quality control resulting in high-quality genotypes, and large sample sizes sufficient in separating out highly correlated signals. There are several different methods to perform fine-mapping, and all methods produce a posterior probability that a variant in that locus is causal. Because the requirements are often difficult to satisfy, there are still limited examples of these methods being more generally applied.
See also
[edit | edit source]- Portal:Biology
- Association mapping
- Epidemiology
- Gene–environment interaction
- Genomics
- Linkage disequilibrium
- Molecular epidemiology
References
[edit | edit source]- ↑ McCarthy, Mark I, ed (October 2010). "Four novel Loci (19q13, 6q24, 12q24, and 5q14) influence the microcirculation in vivo". PLoS Genetics 6 (10): e1001184. doi:10.1371/journal.pgen.1001184. PMID 21060863. PMC 2965750. //www.ncbi.nlm.nih.gov/pmc/articles/PMC2965750/.
- ↑ 2.0 2.1 "Genomewide association studies and assessment of the risk of disease". The New England Journal of Medicine 363 (2): 166–76. July 2010. doi:10.1056/NEJMra0905980. PMID 20647212.
- ↑ 3.0 3.1 3.2 "How to interpret a genome-wide association study". JAMA 299 (11): 1335–44. March 2008. doi:10.1001/jama.299.11.1335. PMID 18349094.
- ↑ "Genome-Wide Association Studies". [[w:National Human Genome Research Institute|]].
- ↑ Ozaki, K (2002). "Functional SNPs in the lymphotoxin-alpha gene that are associated with susceptibility to myocardial infarction". Nature Genetics 19: 212–219. https://www.nature.com/articles/ng1047.
- ↑ "Complement factor H polymorphism in age-related macular degeneration". Science 308 (5720): 385–9. April 2005. doi:10.1126/science.1109557. PMID 15761122. PMC 1512523. //www.ncbi.nlm.nih.gov/pmc/articles/PMC1512523/.
- ↑ "GWAS Catalog: The NHGRI-EBI Catalog of published genome-wide association studies". European Molecular Biology Laboratory. European Molecular Biology Laboratory. Retrieved 18 April 2017.
- ↑ 8.0 8.1 8.2 Lewitter, Fran; Kann, Maricel, eds (2012). "Chapter 11: Genome-wide association studies". PLoS Computational Biology 8 (12): e1002822. doi:10.1371/journal.pcbi.1002822. PMID 23300413. PMC 3531285. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3531285/.
- ↑ 9.0 9.1 Human Molecular Genetics (4th ed.). Garland Science. pp. 467–495. ISBN 978-0-8153-4149-9.
- ↑ "Online Mendelian Inheritance in Man". Archived from the original on 5 December 2011. Retrieved 2011-12-06.
- ↑ "Genomewide scans of complex human diseases: true linkage is hard to find". American Journal of Human Genetics 69 (5): 936–50. November 2001. doi:10.1086/324069. PMID 11565063. PMC 1274370. //www.ncbi.nlm.nih.gov/pmc/articles/PMC1274370/.
- ↑ "The future of genetic studies of complex human diseases". Science 273 (5281): 1516–7. September 1996. doi:10.1126/science.273.5281.1516. PMID 8801636.
- ↑ "The uneasy ethical and legal underpinnings of large-scale genomic biobanks". Annual Review of Genomics and Human Genetics 8: 343–64. 2007. doi:10.1146/annurev.genom.7.080505.115721. PMID 17550341.
- ↑ "The International HapMap Project". Nature 426 (6968): 789–96. December 2003. doi:10.1038/nature02168. PMID 14685227.
- ↑ "Quantitative monitoring of gene expression patterns with a complementary DNA microarray". Science 270 (5235): 467–70. October 1995. doi:10.1126/science.270.5235.467. PMID 7569999.
- ↑ 16.0 16.1 16.2 16.3 Wellcome Trust Case Control Consortium, Burton PR (June 2007). "Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls". Nature 447 (7145): 661–78. doi:10.1038/nature05911. PMID 17554300. PMC 2719288. //www.ncbi.nlm.nih.gov/pmc/articles/PMC2719288/.
- ↑ 17.0 17.1 17.2 17.3 "Basic statistical analysis in genetic case-control studies". Nature Protocols 6 (2): 121–33. February 2011. doi:10.1038/nprot.2010.182. PMID 21293453. PMC 3154648. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3154648/.
- ↑ "PLINK: a tool set for whole-genome association and population-based linkage analyses". American Journal of Human Genetics 81 (3): 559–75. September 2007. doi:10.1086/519795. PMID 17701901. PMC 1950838. //www.ncbi.nlm.nih.gov/pmc/articles/PMC1950838/.
- ↑ "MOBAS: identification of disease-associated protein subnetworks using modularity-based scoring". EURASIP Journal on Bioinformatics & Systems Biology 2015 (1): 7. December 2015. doi:10.1186/s13637-015-0025-6. PMID 28194175. https://link.springer.com/article/10.1186/s13637-015-0025-6.
- ↑ Ayati, Marzieh; Koyutürk, Mehmet (2015-01-01). "Assessing the Collective Disease Association of Multiple Genomic Loci". Proceedings of the 6th ACM Conference on Bioinformatics, Computational Biology and Health Informatics. BCB '15 (New York, NY, USA: ACM): 376–385. doi:10.1145/2808719.2808758. ISBN 978-1-4503-3853-0. http://doi.acm.org/10.1145/2808719.2808758.
- ↑ "Genotype imputation for genome-wide association studies". Nature Reviews. Genetics 11 (7): 499–511. July 2010. doi:10.1038/nrg2796. PMID 20517342.
- ↑ "Genotype imputation with thousands of genomes". G3 1 (6): 457–70. November 2011. doi:10.1534/g3.111.001198. PMID 22384356. PMC 3276165. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3276165/.
- ↑ "[Chronic myelogenous leukaemia as a possible consequence of immunosuppressive treatment of nephrotic syndrome (author's transl)]". Monatsschrift Fur Kinderheilkunde 123 (10): 718–20. October 1975. PMID 1058334.
- ↑ "Genes mirror geography within Europe". Nature 456 (7218): 98–101. November 2008. doi:10.1038/nature07331. PMID 18758442. PMC 2735096. //www.ncbi.nlm.nih.gov/pmc/articles/PMC2735096/.
- ↑ Charney, Evan (2016-12-01). "Genes, behavior, and behavior genetics". Wiley Interdisciplinary Reviews: Cognitive Science 8 (1-2): e1405. doi:10.1002/wcs.1405. ISSN 1939-5078. http://doi.wiley.com/10.1002/wcs.1405.
- ↑ "A novel computational biostatistics approach implies impaired dephosphorylation of growth factor receptors as associated with severity of autism". Translational Psychiatry 4 (1): e354. January 2014. doi:10.1038/tp.2013.124. PMID 24473445. PMC 3905234. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3905234/.
- ↑ "Guidelines for genome-wide association studies". PLoS Genetics 8 (7): e1002812. July 2012. doi:10.1371/journal.pgen.1002812. PMID 22792080. PMC 3390399. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3390399/.
- ↑ Gibson, Greg, ed (July 2011). "Fine mapping of five loci associated with low-density lipoprotein cholesterol detects variants that double the explained heritability". PLoS Genetics 7 (7): e1002198. doi:10.1371/journal.pgen.1002198. PMID 21829380. PMC 3145627. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3145627/.
- ↑ "Potential etiologic and functional implications of genome-wide association loci for human diseases and traits". Proceedings of the National Academy of Sciences of the United States of America 106 (23): 9362–7. June 2009. doi:10.1073/pnas.0903103106. PMID 19474294. PMC 2687147. //www.ncbi.nlm.nih.gov/pmc/articles/PMC2687147/.
- ↑ "An open access database of genome-wide association results". BMC Medical Genetics 10: 6. January 2009. doi:10.1186/1471-2350-10-6. PMID 19161620. PMC 2639349. //www.ncbi.nlm.nih.gov/pmc/articles/PMC2639349/.
- ↑ "Complement factor H variant increases the risk of age-related macular degeneration". Science 308 (5720): 419–21. April 2005. doi:10.1126/science.1110359. PMID 15761120.
- ↑ "Design and development of TT30, a novel C3d-targeted C3/C5 convertase inhibitor for treatment of human complement alternative pathway-mediated diseases". Blood 118 (17): 4705–13. October 2011. doi:10.1182/blood-2011-06-359646. PMID 21860027. PMC 3208285. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3208285/.
- ↑ "Largest ever study of genetics of common diseases published today" (Press release). Wellcome Trust Case Control Consortium. 6 June 2007. Retrieved 19 June 2008.
- ↑ "Validating, augmenting and refining genome-wide association signals". Nature Reviews. Genetics 10 (5): 318–29. May 2009. doi:10.1038/nrg2544. PMID 19373277.
- ↑ "Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals". Nature Genetics 50 (8): 1112 - 1121. August 2018. doi:10.1038/s41588-018-0147-3. PMID 30038396.
- ↑ Genome-wide Analysis of Insomnia (N=1,331,010) Identifies Novel Loci and Functional Pathways. January 2018. doi:10.1101/214973.
- ↑ 37.0 37.1 37.2 "Common variants at 30 loci contribute to polygenic dyslipidemia". Nature Genetics 41 (1): 56–65. January 2009. doi:10.1038/ng.291. PMID 19060906. PMC 2881676. //www.ncbi.nlm.nih.gov/pmc/articles/PMC2881676/.
- ↑ "Genome-wide association identifies nine common variants associated with fasting proinsulin levels and provides new insights into the pathophysiology of type 2 diabetes". Diabetes 60 (10): 2624–34. October 2011. doi:10.2337/db11-0415. PMID 21873549. PMC 3178302. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3178302/.
- ↑ "C-reactive protein and coronary disease: is there a causal link?". Circulation 120 (21): 2036–9. November 2009. doi:10.1161/CIRCULATIONAHA.109.907212. PMID 19901186.
- ↑ "Case-control association mapping by proxy using family history of disease". Nature Genetics 49 (3): 325–331. March 2017. doi:10.1038/ng.3766. PMID 28092683.
- ↑ "The pursuit of genome-wide association studies: where are we now?". Journal of Human Genetics 55 (4): 195–206. April 2010. doi:10.1038/jhg.2010.19. PMID 20300123.
- ↑ 42.0 42.1 "Personal genomes: The case of the missing heritability". Nature 456 (7218): 18–21. November 2008. doi:10.1038/456018a. PMID 18987709.
- ↑ "Genomics: Hepatitis C virus gets personal". Nature 461 (7262): 357–8. September 2009. doi:10.1038/461357a. PMID 19759611.Template:Closed access
- ↑ "Chromosome 9p21 variant predicts mortality after coronary artery bypass graft surgery". Circulation 122 (11 Suppl): S60-5. September 2010. doi:10.1161/CIRCULATIONAHA.109.924233. PMID 20837927. PMC 2943860. //www.ncbi.nlm.nih.gov/pmc/articles/PMC2943860/.
- ↑ "Association between a literature-based genetic risk score and cardiovascular events in women". JAMA 303 (7): 631–7. February 2010. doi:10.1001/jama.2010.119. PMID 20159871. PMC 2845522. //www.ncbi.nlm.nih.gov/pmc/articles/PMC2845522/.
- ↑ 46.0 46.1 "Major heart disease genes prove elusive". Science 328 (5983): 1220–1. June 2010. doi:10.1126/science.328.5983.1220. PMID 20522751.Template:Closed access
- ↑ "Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance". Nature 461 (7262): 399–401. September 2009. doi:10.1038/nature08309. PMID 19684573.
- ↑ "Genetic variation in IL28B and spontaneous clearance of hepatitis C virus". Nature 461 (7265): 798–801. October 2009. doi:10.1038/nature08463. PMID 19759533. PMC 3172006. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3172006/.
- ↑ "Personalized medicine and human genetic diversity". Cold Spring Harbor Perspectives in Medicine 4 (9): a008581. July 2014. doi:10.1101/cshperspect.a008581. PMID 25059740. PMC 4143101. //www.ncbi.nlm.nih.gov/pmc/articles/PMC4143101/.
- ↑ "Association of genetic risk variants with expression of proximal genes identifies novel susceptibility genes for cardiovascular disease". Circulation: Cardiovascular Genetics 3 (4): 365–73. August 2010. doi:10.1161/CIRCGENETICS.110.948935. PMID 20562444.
- ↑ "Abdominal aortic aneurysm is associated with a variant in low-density lipoprotein receptor-related protein 1". American Journal of Human Genetics 89 (5): 619–27. November 2011. doi:10.1016/j.ajhg.2011.10.002. PMID 22055160. PMC 3213391. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3213391/.
- ↑ "A genome-wide association study in Europeans and South Asians identifies five new loci for coronary artery disease". Nature Genetics 43 (4): 339–44. March 2011. doi:10.1038/ng.782. PMID 21378988. PMC 3190399. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3190399/.Template:Closed access
- ↑ "Blood pressure loci identified with a gene-centric array". American Journal of Human Genetics 89 (6): 688–700. December 2011. doi:10.1016/j.ajhg.2011.10.013. PMID 22100073. PMC 3234370. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3234370/.
- ↑ "Sortilin: an unusual suspect in cholesterol metabolism: from GWAS identification to in vivo biochemical analyses, sortilin has been identified as a novel mediator of human lipoprotein metabolism". BioEssays 33 (6): 430–7. June 2011. doi:10.1002/bies.201100003. PMID 21462369.Template:Closed access
- ↑ "Functional validation of new pathways in lipoprotein metabolism identified by human genetics". Current Opinion in Lipidology 22 (2): 123–8. April 2011. doi:10.1097/MOL.0b013e32834469b3. PMID 21311327.Template:Closed access
- ↑ Pickrell J, Barrett J, MacArthur D, Jostins L (23 November 2011). "Size matters, and other lessons from medical genetics". Genomes Unzipped. Retrieved 7 December 2011.
- ↑ "Genetic signatures of exceptional longevity in humans". Science 2010. July 2010. doi:10.1126/science.1190532. PMID 20595579.Template:RetractedTemplate:Closed access
- ↑ MacArthur, Daniel. "Serious flaws revealed in "longevity genes" study". Wired. https://www.wired.com/wiredscience/2010/07/serious-flaws-revealed-in-longevity-genes-study/. Retrieved 2011-12-07.
- ↑ "Retraction". Science 333 (6041): 404. July 2011. doi:10.1126/science.333.6041.404-a. PMID 21778381.Template:Closed access
- ↑ "Genetic signatures of exceptional longevity in humans". PLOS One 7 (1): e29848. 2012-01-18. doi:10.1371/journal.pone.0029848. PMID 22279548. PMC 3261167. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3261167/.
- ↑ "Five years of GWAS discovery". American Journal of Human Genetics 90 (1): 7–24. January 2012. doi:10.1016/j.ajhg.2011.11.029. PMID 22243964. PMC 3257326. //www.ncbi.nlm.nih.gov/pmc/articles/PMC3257326/.
- ↑ "Evidence-based psychiatric genetics, AKA the false dichotomy between common and rare variant hypotheses". Molecular Psychiatry 17 (5): 474–85. May 2012. doi:10.1038/mp.2011.65. PMID 21670730.Template:Closed access
External links
[edit | edit source]- Genotype-phenotype interaction software tools and databases on Omictools
- Statistical Methods for the Analysis of Genome-Wide Association Studies [video lecture series]
- Whole genome association studies — by the National Human Genome Research Institute
- GWAS Central — a central database of summary-level genetic association findings
- Barrett, Jeff (18 July 2010). "How to read a genome-wide association study". Genomes Unzipped.
- Consortia of genome-wide association studies (GWAS) — by Bennett SN, Caporaso, NE, et al.
- PLINK — whole genome association analysis toolset
- ENCODE threads explorer Impact of functional information on understanding variation. Nature
- Custom Genome-Wide Association Studies

